28 August 2020: Clinical Research
Circ_cse1l Inhibits Colorectal Cancer Proliferation by Binding to eIF4A3
Bin Xu1ABCDEF, Ning Yang1BCDF, Yabin Liu1BCDF, Peng Kong2ACE, Mei Han2AG, Binghui Li1ABCDEFG*DOI: 10.12659/MSM.923876
Med Sci Monit 2020; 26:e923876
Abstract
BACKGROUND: Circular RNAs (circRNAs) are involved in the growth of many tumors. However, the expression and possible role of circ_cse1l (hsa_circ_0060745) in colorectal cancer (CRC) are unclear. The present study was designed to explore the role of circ_cse1l in CRC.
MATERIAL AND METHODS: The levels of circ_cse1l expression in cancer tissues and serum samples of 50 patients with CRC and in control subjects were analyzed by quantitative reverse transcription-polymerase chain reaction (qRT-PCR). CCK-8, colony formation, transwell and wound healing assays were performed to assess the functions of circ_cse1l in CRC cell lines after overexpression. The relationship between circ_cse1l and eIF4A3 during cell proliferation was analyzed by western blotting and RNA-binding protein immunoprecipitation (RIP).
RESULTS: qRT-PCR assays showed that the levels of expression of circ_cse1l were lower in CRC cell lines and in tissue and serum samples from patients with CRC than in control samples. The expression of circ_cse11 in CRC tissues had clinical significance, as its level of expression was inversely associated with the depth of tumor invasion. Overexpression of circ_cse1l in HT29 and HCT116 cells markedly reduced cell proliferation and metastasis. Western blotting showed that circ_cse1l overexpression dowregulated the expression of PCNA protein. RIP results demonstrated that circ_cse1l inhibited the proliferation of CRC cells by binding to eIF4A3.
CONCLUSIONS: The expression of circ_cse1l is downregulated in CRC. Furthermore, circ_cse1l downregulated PCNA expression by binding to eIF4A3, inhibiting the proliferation of CRC cells.
Keywords: Biological Markers, Colorectal Neoplasms, Diagnosis, DNA, Circular, Eukaryotic Initiation Factor-4A, Aged, 80 and over, DEAD-box RNA Helicases, Down-Regulation, HCT116 Cells, HT29 Cells, RNA, Circular, Real-Time Polymerase Chain Reaction, Reverse Transcriptase Polymerase Chain Reaction
Background
Colorectal cancer (CRC) is the third most common type of cancer worldwide [1], with nearly 2 million persons newly diagnosed with CRC and nearly 900,000 patients dying of this disease in 2018 [2]. The rate of distant metastasis in CRC patients is about 40%, and the recurrence rate in the absence of lymph node metastasis is about 25% [3,4]. Despite improvements in its diagnosis and treatment and the elucidation of some of the molecular mechanisms of CRC, CRC incidence and mortality rates remain high [5]. At present, the molecular markers commonly used in the clinical diagnosis of CRC include carcinoembryonic antigen (CEA), CA199 and markers of inflammation such as NLR and PLR. These markers, however, are relatively insensitive and are therefore used as markers of tumor recurrence [6–8]. Therefore, it is necessary to identify biomarkers effective in the diagnosis of CRC.
Circular RNA (circRNA) is a series of functional molecules involved in many physiological and pathological processes. CircRNAs are mainly formed by RNA splicing at the 5′ end of the upstream exon and the 3′ end of the downstream exon [9]. Compared with conventional linear RNA, the structure of circular RNA is more stable [10]. Moreover, circRNAs may be potential biomarkers in tumors. Studies have shown that several circRNAs, including Circ_0104631 and circZNF609, are abnormally expressed in CRC and may have biological functions in these tumors [11–12]. However, the biological function of most circular RNAs in CRC is still unclear.
CSE1L, originally isolated from breast cancer cells [13], has been associated with the invasion, metastasis and proliferation of many tumors, including bladder [14], lung [15], breast [16] and colorectal [17] cancers. Knock-down of CSE1L was found to inhibit the expression of CRC cells and promote their apoptosis [18]. The biological properties of the circRNA derived from CSE1L has not yet been assessed in CRC. The present study assessed the expression, function and mechanism of action of circ_cse1l in CRC.
Material and Methods
PATIENTS AND SAMPLE COLLECTION:
The study was approved by the ethics committee of our institution, and informed consent was obtained from each patient. Between March 2018 and July 2018, tissue and serum samples were obtained from 50 CRC patients diagnosed and treated at our center. Patients were included if they were pathologically confirmed as having primary CRC, if radical resection was feasible, if their clinicopathological data were complete, and if they had not received radiotherapy, chemotherapy or any other neoadjuvant therapy before surgery. Tissue and serum samples were obtained within 20 minutes after surgery and stored in an ultra-low temperature freezer. As controls, peripheral blood samples were obtained from 50 healthy individuals. Clinical characteristics of all patients were recorded.
CELL CULTURE AND TRANSFECTION:
The FHC, HT29, HCT116, and LoVo cell lines were obtained from the American Type Culture Collection (ATCC; Manassas, VA, USA). FHC cells were cultured in complete DMEM (Invitrogen, USA), and the other cell lines were cultured in McCoy’s 5A complete medium containing 10% serum. All cells were cultured in a CO2 incubator at constant temperature, and the medium was changed regularly. The circ_cse1l overexpression vector was constructed by Sangon Biotech (Beijing, China), with the empty pCDNA3.1 vector being a negative control. HT29 and HCT116 cells were transfected with these vectors at a certain cell density.
RNA EXTRACTION AND QUANTITATIVE REAL-TIME POLYMERASE CHAIN REACTION (QRT-PCR):
Total RNA was extracted from samples using TRIzol reagent (Life Technologies, USA) and cDNA was synthesized from RNA by reverse transcription using M-MLV First Strand Kits (Beyotime, Beijing, China). qRT-PCR assays were performed using SYBR Green qPCR SuperMix-UDG (Beyotime) and primers for
Levels of expression of circ_cse1l and PCNA were normalized to those of GAPDH using the 2−ΔΔCT method.
CELL COUNTING KIT-8 (CCK-8) ASSAY:
Each well of a 96-well plate was seeded with about 5000 tumor cells. The cells were transfected with circ_cse1l overexpression vector or empty pCDNA3.1 vector, and cell growth was measured 24, 48, 72, and 96 h later by the CCK-8 method (APExBIO, Houston, USA). Briefly 10 μl CCK-8 reagents were added to each well, the plates were incubated at 37°C for 2 hours, and the absorbance of cells in each well was measured at 450 nm using a microplate reader.
COLONY FORMATION ASSAY:
Transfected, logarithmically growing cells were cultured for 24 hours. Each well of a six-well plate was seeded with 500 cells, and the plates were cultured for 2 weeks. The cells were subsequently washed, fixed, stained, and dried. The numbers of colonies in each well were counted and photographed.
WOUND HEALING ASSAY:
Before the experiment, a horizontal line was drawn evenly behind the six-well plate. Twenty-four hours after transfection, tumor cells were plated uniformly in six-well plates. The cells adherent to each well were scratched with a 200 μl pipette tip, and each well was washed twice with PBS to remove the detached cells. The remaining cells were cultured for 24 hours, and the area of each scratch was assessed with a microscope.
TRANSWELL INVASION ASSAY:
The invasive ability of the cells was detected using transwell chambers. Briefly, 50 μl of Matrigel (Invitrogen, USA) were spread onto the upper surface of each membrane, and the 24-well plate was cultured for 6 hours. Twenty-four hours after transfection, HT29 and HCT116 cells were starved in serum-free medium for 24 hours. After adjusting the cell concentration, 1×105 cells and 200 μl serum-free medium were added to the upper chamber of each well, and an appropriate amount of McCoy’s 5A complete medium was added to the lower chamber of each Transwell. After culture for 24 hours, the cells in the upper chamber were gently wiped with a cotton swab. Finally, the invasive cells were fixed for 20 min, stained and counted under a microscope.
WESTERN BLOTTING:
HT29 and HCT116 cells were transfected with vector and incubated for 48 hours. Total proteins were extracted using lysis buffer, and the total protein concentration in each lysate was measured using the Lowry method. Aliquots of 30 μg protein were loaded onto each well of an SDS-PAGE gel. Following electrophoresis, the proteins were transferred to a PVDF membrane. The membranes were incubated for 2 hours in solution containing 5% skim milk to block nonspecific binding and Incubated with a 1: 500 dilution of primary anti-PCNA antibody (Abcam, USA) or a 1: 1000 dilution of primary anti-GAPDH antibody (Abcam, USA) at 4°C overnight with gentle shaking. After washing, the PVDF membranes were incubated with diluted secondary antibody solution for 1 hour. The membranes were scanned using a chemiluminescence imaging analyzer.
RNA IMMUNOPRECIPITATION ASSAY (RIP):
RIP experiments were performed using RIP kits, according to the protocol described by the manufacturer (Millipore, USA). Cells were lysed and their lysates were cleared and incubated with magnetic beads and anti-eIF4A3 antibodies or with non-specific IgG as a control. The beads were washed for RNA extraction, and the enriched RNA was analyzed by qRT-PCR.
STATISTICAL ANALYSIS:
All data were analyzed using SPSS software 23.0 (IBM, USA). The expression of circ_cse11 in two groups was compared by t-tests, and the clinical significance of circ_cse11 expression in CRC was analyzed by independent sample t-tests. A P-value <0.05 was defined as statistically significant.
Results
PATIENTS:
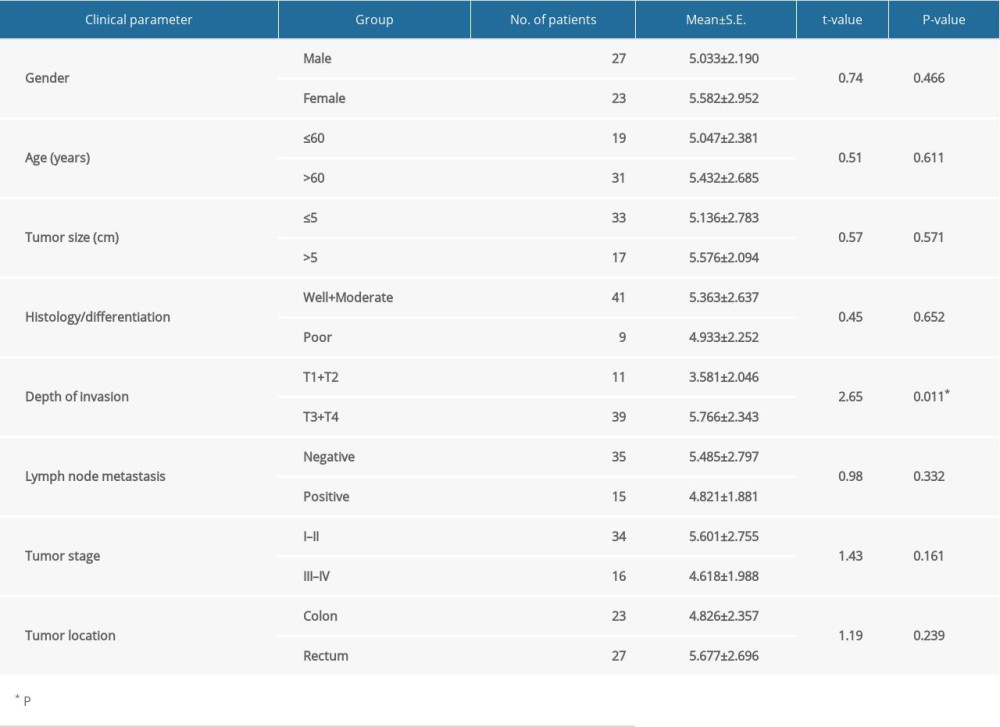
This study included 50 patients, 27 men and 23 women, of median age 60 years (range 30–86 years). Of these 50 patients, 27 had rectal cancer and 23 had colon cancer. Postoperative pathological data showed that 33 patients had tumors ≤5 cm in diameter, whereas 17 had tumors >5 cm. The tumors of 41 patients were well/moderately differentiated, whereas the other nine tumors were poorly differentiated. According to TNM staging criteria, 11 patients were T1+T2 and 39 were T3+T4; 15 patients had lymph node metastasis, whereas 35 did not; and 34 patients were stage I–II and 16 were stage III–IV.
CIRC_CSE1L IS DOWNREGULATED IN CRC AND RELATED TO CLINICAL CHARACTERISTICS:
Of the 12 circRNAs from the host gene cse1l detected in CRC samples (Figure 1A), we chose the circ_cse1l located at chr20: 47688822–47689234 in the human genome (hsa_circ_0060745) as our research target. Expression of circ_cse1l was significantly lower in tissues and serum samples of the 50 CRC patients than in normal controls group (P<0.05) (Figure 1B). In addition, circ_cse1l was expressed in FHC cells and CRC cell lines (HT29, HCT116, LoVo), and we tested the effect of circ_cse1l overexpression in HT29 and HCT116 cells (Figure 1C).
Investigation of the clinical significance of circ_cse11 expression in CRC showed that its level of expression was inversely associated with the depth of tumor invasion (Table 1).
OVEREXPRESSION OF CIRC_CSE1L INHIBITS THE ABILITY OF CRC CELLS TO MIGRATE AND INVADE:
We evaluated the effect of circ_cse1l expression on the ability of CRC cells to migrate and invade other tissues. Wound healing experiments showed that the overexpression of circ_cse1l reduced the ability of HT29 and HCT116 cells to migrate (Figure 2A), whereas Transwell invasion experiments showed that circ_cse1l expression reduced their invasive ability (Figure 2B).
OVEREXPRESSION OF CIRC_CSE1L INHIBITS CRC CELL PROLIFERATION:
CCK-8 assessment of the effect of circ_cse1l expression on CRC cell proliferation showed that circ_cse1l overexpression inhibited the proliferation of HT29 and HCT116 cells (Figure 3A). Similar findings were observed in colony formation experiments (Figure 3B). The proliferation marker PCNA is closely associated with cell DNA synthesis and the initiation of the cell proliferation process, an indicator of the development of CRC [19]. Therefore, we assessed the effects of circ_cse1l overexpression on PCNA protein expression in HT29 and HCT116 cells, finding that circ_cse1l overexpression reduced the expression of PCNA (Figure 3C).
CIRC_CSE1L CAN CONTROL PCNA EXPRESSION BY BINDING TO EIF4A3:
Previous studies have found that, in addition to acting as a miRNA sponge-binding RNA, circRNA can bind to proteins to exert regulatory effects [20]. So, we used CircInteractome (https://circinteractome.nia.nih.gov/) to predict proteins that potentially bind to circ_cse1l. The RIP experiment results identified eIF4A3 as binding to circ_cse1l (Figure 4A). EIF4A3 is a eukaryotic translation initiation factor that can monitor the quality of mRNA before it enters a translation event. Using the RIP method, we found that the level of PCNA mRNA in fraction immunoprecipitated by anti-eIF4A3 was reduced when circ_cse1l was overexpressed (Figure 4B). These findings indicate that circ_cse11 reduces PCNA expression levels by binding to eIF4A3.
Discussion
CRC patients are widely distributed worldwide. Despite recent progress in treatment methods, tumor recurrence and metastasis may still occur in 40% to 50% of patients, and their prognosis is poor [21]. Therefore, it is necessary to identify sensitive biomarkers for the diagnosis of CRC. CircRNA is a special type of endogenous non-coding RNA (ncRNA) [22]. circRNA is differentially expressed in various types of malignant tumor, including CRC, with its level of expression being closely related to clinical stage and prognosis [23,24].
CSE1L is highly expressed in CRC and plays important roles in tumor proliferation, invasion, migration and apoptosis [25]. The present study showed that circ_cse11 derived from the
PCNA is a marker of cell proliferation, especially in tumors. The level of expression of PCNA was found to be related to the degree of malignancy of CRC, distant metastasis and patient survival rate, and can be used as a biomarker for CRC proliferation [26]. Circ_0137008 was shown to reduce the viability and colony-forming ability of CRC cells by inhibiting PCNA expression [27]. In exploring the relationship between circ_cse11 and PCNA, we found that overexpression of circ_cse11 inhibited the expression of PCNA, suggesting that circ_cse11 inhibits the proliferation of CRC cells by down-regulating the level of expression of PCNA.
CircRNA can not only bind miRNA and act as a “miRNA sponge”, but can also directly regulate molecular mechanisms through interaction with RNA-binding proteins, especially in tumors. Overexpression of circ-Foxo3 resulted in a greater degree of binding HIF1α and FAK, inhibiting their function and accelerating the process of cell aging [28]. Circ_0014717 was found to inhibit CRC tumors by affecting p16 protein expression [29]. Moreover, circ-0075804 was found to promote the proliferation of retinoblastoma cells by combining with HNRNPK to improve the stability of E2F3 [30]. Using CircInteractome, we predicted that eIF4A3 was a protein that could bind to circ_cse1l and confirmed this binding of eIF4A3 to circ_cse1l by RIP experiments.
The EJC formed by eIF4A3 can trigger meaningless mediated mRNA decay, affecting mRNA levels and regulating protein expression at the translational and post-translational levels [31,32]. In addition, eIF4A3 has been shown important in certain cancers. For example, circPVRL3 was found to promote the proliferation of gastric cancer cells by combining with eIF4A3 [33]. CircSEPT9, together with E2F1 and EIF4A3, has been shown promote the occurrence and development of breast cancer through the circSEPT9/miR-637/LIF axis [34]. In addition, eIF4A3 has been reported to affect mRNA expression at the post-transcriptional level, thereby affecting the biological behavior of tumors. Long noncoding RNA H19 was found to bind to eIF4A3, thereby regulating the levels of expression of mRNAs encoding cell cycle-related factors, such as cyclin D1, cyclin E1 and CDK4, and thereby affecting the proliferation of CRC cells [35].
Our experimental results suggest that circ_cse1l binding of eIF4A3 affects the level of expression of PCNA mRNA, a finding confirmed by RIP experiments. These results are consistent previous findings [35], indicating that eIF4A3 can regulate PCNA expression at the mRNA level. To our knowledge, this study is the first to show that circ_cse1l can down-regulate the expression of PCNA by binding to eIF4A3, which in turn affected the proliferation of CRC cells. Additional studies, however, are required to further clarify the role of circ_cse1l in CRC.
Conclusions
This study showed that the expression of circ_cse11 was downregulated in CRC and that its level of expression was inversely related to the depth of tumor invasion. These results also showed that circ_cse1l could downregulate PCNA expression by binding to eIF4A3, thereby inhibiting the proliferation of CRC cells.
Figures
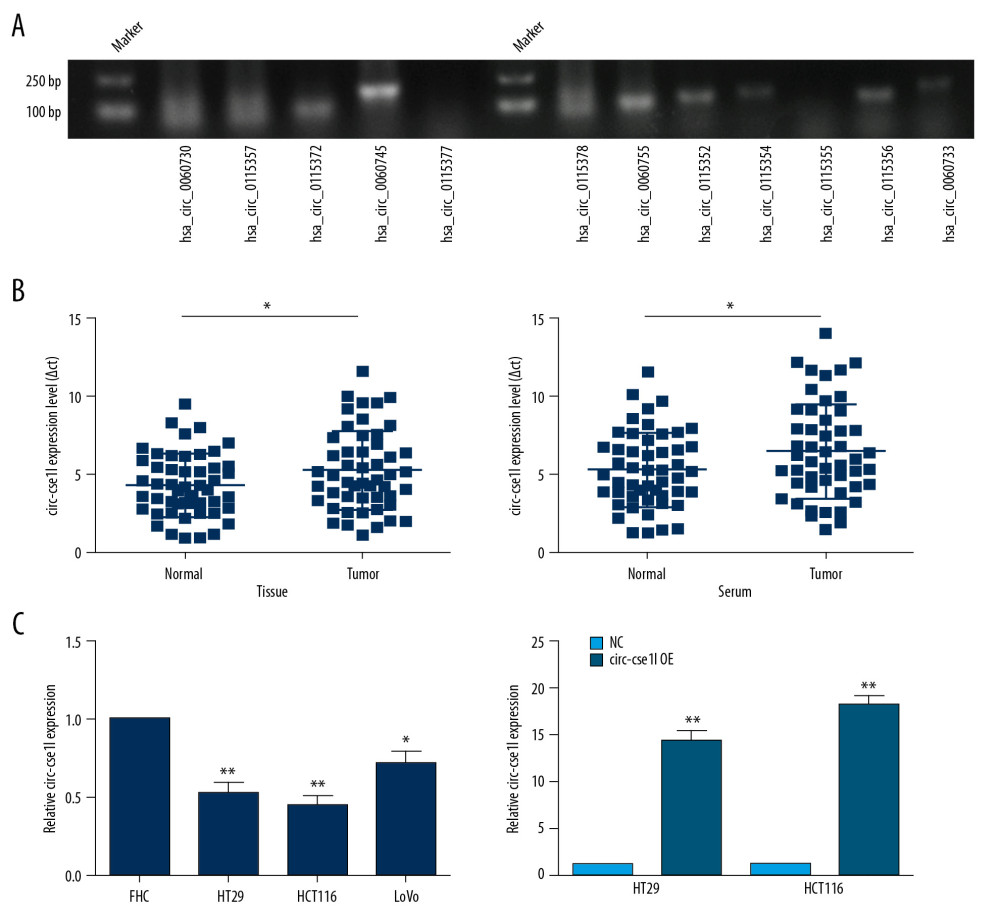 Figure 1. Downregulation of circ_cse1l expression in CRC, as shown by qRT-PCR. (A) Expression in CRC of 12 circRNAs encoded by the CSE1L gene. (B) Relative expression of circ_cse1l in tissue and serum samples from patients with CRC and normal controls. (C) Expression of circ_cse1l in FHC, HT29, HCT116, and LoVo cells and after transfection of the circ_cse1l overexpression vector and empty vector (NC) into HT29 and HCT116 cells. * P<0.05, ** P<0.01.
Figure 1. Downregulation of circ_cse1l expression in CRC, as shown by qRT-PCR. (A) Expression in CRC of 12 circRNAs encoded by the CSE1L gene. (B) Relative expression of circ_cse1l in tissue and serum samples from patients with CRC and normal controls. (C) Expression of circ_cse1l in FHC, HT29, HCT116, and LoVo cells and after transfection of the circ_cse1l overexpression vector and empty vector (NC) into HT29 and HCT116 cells. * P<0.05, ** P<0.01. 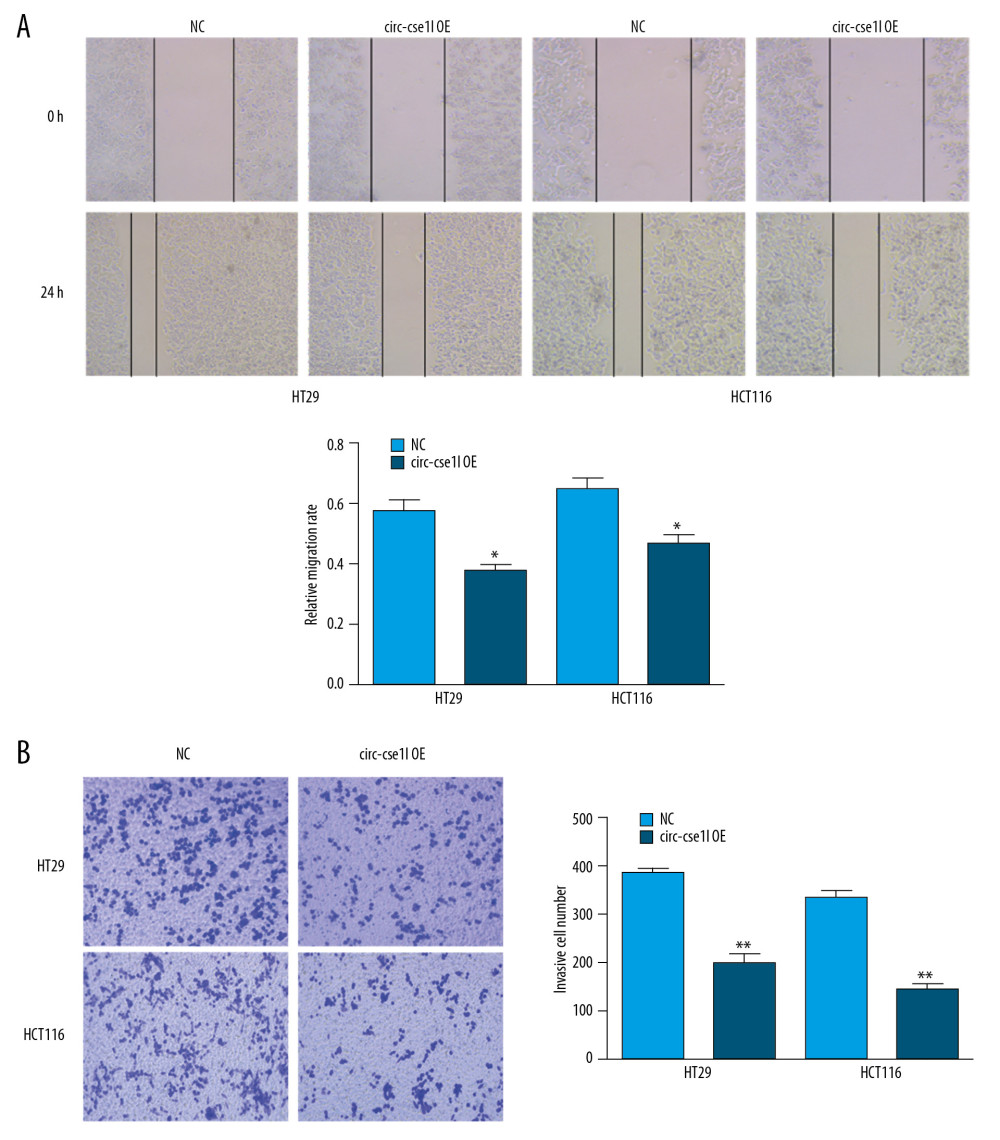 Figure 2. Overexpression of circ_cse1l inhibits the ability of CRC cells to migrate and invade. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). Overexpression of circ_cse1l reduced the (A) migration and (B) invasion of HT29 and HCT116 cells. *P<0.05, ** P<0.01.
Figure 2. Overexpression of circ_cse1l inhibits the ability of CRC cells to migrate and invade. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). Overexpression of circ_cse1l reduced the (A) migration and (B) invasion of HT29 and HCT116 cells. *P<0.05, ** P<0.01. 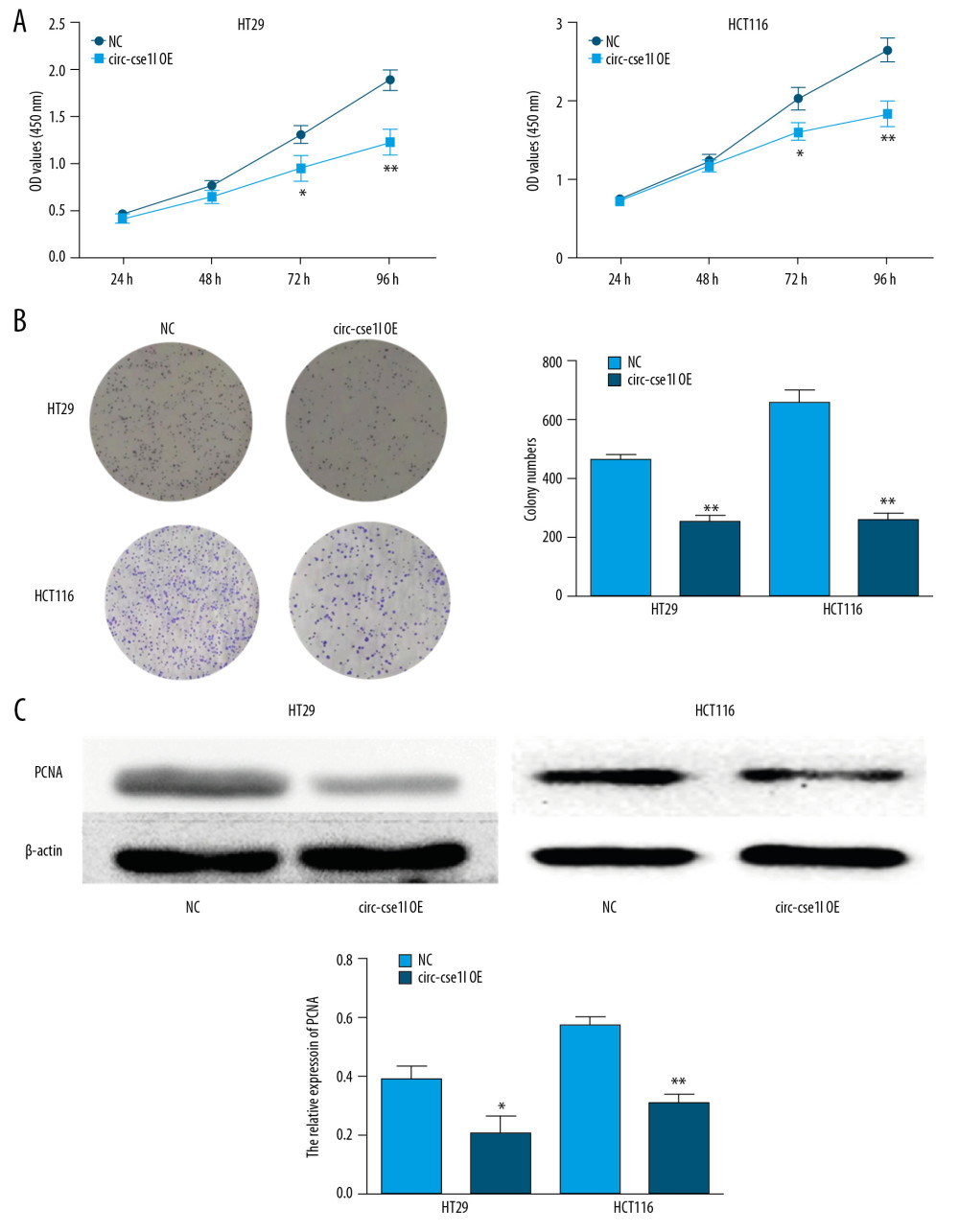 Figure 3. Overexpression of circ_cse1l inhibits CRC cell proliferation while reducing PCNA expression. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). (A) CCK-8 and (B) colony formation assays, showing that overexpression of circ_cse1l inhibited the proliferation of HT29 and HCT116 cells. (C) Western blotting experiments, showing that circ_cse1l overexpression reduced the expression of PCNA protein in HT29 and HCT116 cells. * P<0.05, ** P<0.01.
Figure 3. Overexpression of circ_cse1l inhibits CRC cell proliferation while reducing PCNA expression. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). (A) CCK-8 and (B) colony formation assays, showing that overexpression of circ_cse1l inhibited the proliferation of HT29 and HCT116 cells. (C) Western blotting experiments, showing that circ_cse1l overexpression reduced the expression of PCNA protein in HT29 and HCT116 cells. * P<0.05, ** P<0.01. 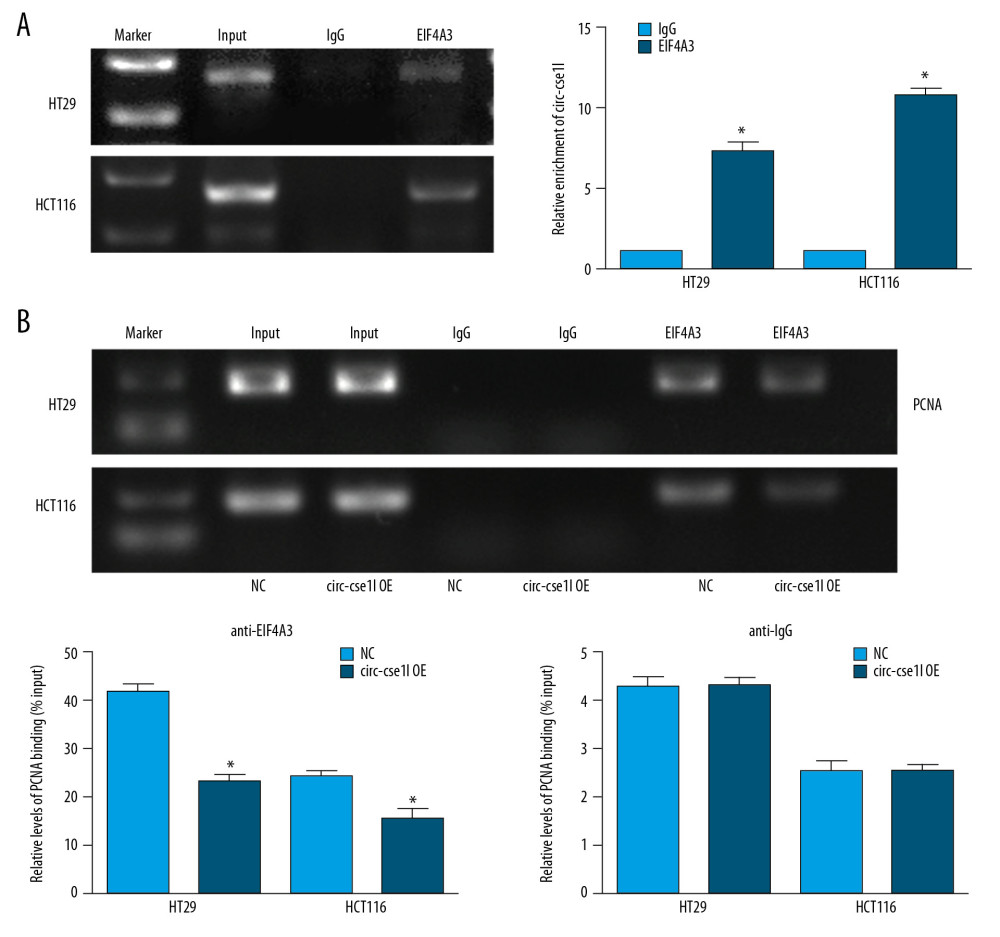 Figure 4. Circ_CSE1L controls PCNA expression by binding to eIF4A3. RIP experiments, showing (A) that eIF4A3 binds to circ_cse1l in HT29 cells and HCT116 cells and (B) that the expression of PCNA mRNA immunoprecipitated by anti-eIF4A3 was reduced by circ_cse1l overexpression. * P<0.05.
Figure 4. Circ_CSE1L controls PCNA expression by binding to eIF4A3. RIP experiments, showing (A) that eIF4A3 binds to circ_cse1l in HT29 cells and HCT116 cells and (B) that the expression of PCNA mRNA immunoprecipitated by anti-eIF4A3 was reduced by circ_cse1l overexpression. * P<0.05. References
1. Siegel RL, Miller KD, Jemal A, Cancer statistics, 2019: Cancer J Clin, 2019; 69; 7-34
2. Bray F, Ferlay J, Soerjomataram I, Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries: Cancer J Clin, 2018; 68; 394-424
3. Ma J, Sun X, Guo T, Interleukin-1 receptor antagonist inhibits angiogenesis via blockage IL-1α/PI3K/NF-κβ pathway in human colon cancer cell: Cancer Manag Res, 2017; 9; 481-93
4. Miao Z, Wang K, Wang X, TNF-α-308G/A polymorphism and the risk of colorectal cancer: A systematic review and an updated meta-analysis: J BUON, 2018; 23; 1616-24
5. Loree JM, Kopetz S, Recent developments in the treatment of metastatic colorectal cancer: Ther Adv Med Oncol, 2017; 9; 551-64
6. Ojima T, Iwahashi M, Nakamura M, Successful cancer vaccine therapy for carcinoembryonic antigen (CEA)-expressing colon cancer using genetically modified dendritic cells that express CEA and T helper-type 1 cytokines in CEA transgenic mice: Int J Cancer, 2007; 120; 585-93
7. Sturgeon CM, Duffy MJ, Stenman UH, National Academy of Clinical Biochemistry laboratory medicine practice guidelines for use of tumor markers in testicular, prostate, colorectal, breast, and ovarian cancers: Clin Chem, 2008; 54; e11-79
8. Li Y, Jia H, Yu W, Nomograms for predicting prognostic value of inflammatory biomarkers in colorectal cancer patients after radical resection: Int J Cancer, 2016; 139; 220-31
9. Zhang Y, Liang W, Zhang P, Circular RNAs: emerging cancer biomarkers and targets: J Exp Clin Cancer Res, 2017; 36; 152
10. Li Y, Zheng Q, Bao C, Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis: Cell Res, 2015; 25; 981-84
11. Cao JZ, Ma LM, Zhang YL, Circ-0104631 promotes cell proliferation and invasion in colorectal cancer and predicts poor prognosis: Eur Rev Med Pharmacol Sci, 2019; 23; 4730-37
12. Zhang X, Zhao Y, Kong P, Expression of circZNF609 is down-regulated in colorectal cancer tissue and promotes apoptosis in colorectal cancer cells by upregulating p53: Med Sci Monit, 2019; 25; 5977-85
13. Brinkmann U, Brinkmann E, Gallo M, Pastan I, Cloning and characterization of a cellular apoptosis susceptibility gene, the human homologue to the yeast chromosome segregation gene CSE1: Proc Natl Acad Sci USA, 1995; 92; 10427-31
14. Chang CC, Tai CJ, Su TC, The prognostic significance of nuclear CSE1L in urinary bladder urothelial carcinomas: Ann Diagn Pathol, 2012; 16; 362-68
15. Papay J, Krenacs T, Moldvay J, Immunophenotypic profiling of nonsmall cell lung cancer progression using the tissue microarray approach: Appl Immunohistochem Mol Morphol, 2007; 15; 19-30
16. Behrens P, Brinkmann U, Fogt F, Implication of the proliferation and apoptosis associated CSE1L/CAS gene for breast cancer development: Anticancer Res, 2001; 21; 2413-17
17. Alnabulsi A, Agouni A, Mitra S, Cellular apoptosis susceptibility (chromosome segregation 1-like, CSE1L) gene is a key regulator of apoptosis, migration and invasion in colorectal cancer: J Pathol, 2012; 228; 471-81
18. Pimiento JM, Neill KG, Henderson-Jackson E: Am J Pathol, 2016; 186; 2761-68
19. Tsai WC, Yu TY, Lin LP, Platelet rich plasma releasate promotes proliferation of skeletal muscle cells in association with upregulation of PCNA, cyclins and cyclin dependent kinases: Platelets, 2017; 28; 491-97
20. Memczak S, Jens M, Elefsinioti A, Circular RNAs are a large class of animal RNAs with regulatory potency: Nature, 2013; 495; 333-38
21. Griffin-Sobel JP, Symptom management of advanced colorectal cancer: Surg Oncol Clin N Am, 2006; 15; 213-22
22. Mercer TR, Dinger ME, Mattick JS, Long non-coding RNAs: insights into functions: Nat Rev Genet, 2009; 10; 155-59
23. Li L, Wan K, Xiong L, CircRNA hsa_circ_0087862 acts as an oncogene in non-small cell lung cancer by targeting miR-1253/RAB3D axis: Onco Targets Ther, 2020; 13; 2873-86
24. Tu FL, Guo XQ, Wu HX, Circ-0001313/miRNA-510-5p/AKT2 axis promotes the development and progression of colon cancer: Am J Transl Res, 2020; 12; 281-91
25. Zhu JH, Hong DF, Song YM, Suppression of cellular apoptosis susceptibility (CSE1L) inhibits proliferation and induces apoptosis in colorectal cancer cells: Asian Pac J Cancer Prev, 2013; 14; 1017-21
26. Yang HB, Hsu PI, Chan SH, Growth kinetics of colorectal adenoma-carcinoma sequence: An immunohistochemical study of proliferating cell nuclear antigen expression: Hum Pathol, 1996; 27; 1071-76
27. Yang Z, Zhang J, Lu D, Hsa_circ_0137008 suppresses the malignant phenotype in colorectal cancer by acting as a microRNA-338-5p sponge: Cancer Cell Int, 2020; 20; 67
28. Du WW, Yang W, Chen Y, Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses: Eur Heart J, 2017; 38; 1402-12
29. Wang F, Wang J, Cao X, Hsa_circ_0014717 is downregulated in colorectal cancer and inhibits tumor growth by promoting p16 expression: Biomed Pharmacother, 2018; 98; 775-82
30. Zhao W, Wang S, Qin T, Wang W, Circular RNA (circ-0075804) promotes the proliferation of retinoblastoma via combining heterogeneous nuclear ribonucleoprotein K (HNRNPK) to improve the stability of E2F transcription factor 3 E2F3: J Cell Biochem, 2020; 121; 3516-25
31. Ferraiuolo MA, Lee CS, Ler LW, A nuclear translation-like factor eIF4AIII is recruited to the mRNA during splicing and functions in nonsense-mediated decay: Proc Natl Acad Sci USA, 2004; 101; 4118-23
32. Conti E, Izaurralde E, Nonsense-mediated mRNA decay: Molecular insights and mechanistic variations across species: Curr Opin Cell Biol, 2005; 17; 316-25
33. Sun HD, Xu ZP, Sun ZQ, Down-regulation of circPVRL3 promotes the proliferation and migration of gastric cancer cells: Sci Rep, 2018; 8; 10111
34. Zheng X, Huang M, Xing L, The circRNA circSEPT9 mediated by E2F1 and EIF4A3 facilitates the carcinogenesis and development of triple-negative breast cancer: Mol Cancer, 2020; 19; 73
35. Han D, Gao X, Wang M, Long noncoding RNA H19 indicates a poor prognosis of colorectal cancer and promotes tumor growth by recruiting and binding to eIF4A3: Oncotarget, 2016; 7; 22159-73
Figures
 Figure 1. Downregulation of circ_cse1l expression in CRC, as shown by qRT-PCR. (A) Expression in CRC of 12 circRNAs encoded by the CSE1L gene. (B) Relative expression of circ_cse1l in tissue and serum samples from patients with CRC and normal controls. (C) Expression of circ_cse1l in FHC, HT29, HCT116, and LoVo cells and after transfection of the circ_cse1l overexpression vector and empty vector (NC) into HT29 and HCT116 cells. * P<0.05, ** P<0.01.
Figure 1. Downregulation of circ_cse1l expression in CRC, as shown by qRT-PCR. (A) Expression in CRC of 12 circRNAs encoded by the CSE1L gene. (B) Relative expression of circ_cse1l in tissue and serum samples from patients with CRC and normal controls. (C) Expression of circ_cse1l in FHC, HT29, HCT116, and LoVo cells and after transfection of the circ_cse1l overexpression vector and empty vector (NC) into HT29 and HCT116 cells. * P<0.05, ** P<0.01. Figure 2. Overexpression of circ_cse1l inhibits the ability of CRC cells to migrate and invade. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). Overexpression of circ_cse1l reduced the (A) migration and (B) invasion of HT29 and HCT116 cells. *P<0.05, ** P<0.01.
Figure 2. Overexpression of circ_cse1l inhibits the ability of CRC cells to migrate and invade. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). Overexpression of circ_cse1l reduced the (A) migration and (B) invasion of HT29 and HCT116 cells. *P<0.05, ** P<0.01. Figure 3. Overexpression of circ_cse1l inhibits CRC cell proliferation while reducing PCNA expression. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). (A) CCK-8 and (B) colony formation assays, showing that overexpression of circ_cse1l inhibited the proliferation of HT29 and HCT116 cells. (C) Western blotting experiments, showing that circ_cse1l overexpression reduced the expression of PCNA protein in HT29 and HCT116 cells. * P<0.05, ** P<0.01.
Figure 3. Overexpression of circ_cse1l inhibits CRC cell proliferation while reducing PCNA expression. HT29 and HCT116 cells were transfected with a circ_cse1l overexpression vector and an empty vector (NC). (A) CCK-8 and (B) colony formation assays, showing that overexpression of circ_cse1l inhibited the proliferation of HT29 and HCT116 cells. (C) Western blotting experiments, showing that circ_cse1l overexpression reduced the expression of PCNA protein in HT29 and HCT116 cells. * P<0.05, ** P<0.01. Figure 4. Circ_CSE1L controls PCNA expression by binding to eIF4A3. RIP experiments, showing (A) that eIF4A3 binds to circ_cse1l in HT29 cells and HCT116 cells and (B) that the expression of PCNA mRNA immunoprecipitated by anti-eIF4A3 was reduced by circ_cse1l overexpression. * P<0.05.
Figure 4. Circ_CSE1L controls PCNA expression by binding to eIF4A3. RIP experiments, showing (A) that eIF4A3 binds to circ_cse1l in HT29 cells and HCT116 cells and (B) that the expression of PCNA mRNA immunoprecipitated by anti-eIF4A3 was reduced by circ_cse1l overexpression. * P<0.05. In Press
08 Mar 2024 : Clinical Research
Evaluation of Foot Structure in Preschool Children Based on Body MassMed Sci Monit In Press; DOI: 10.12659/MSM.943765
15 Apr 2024 : Laboratory Research
The Role of Copper-Induced M2 Macrophage Polarization in Protecting Cartilage Matrix in OsteoarthritisMed Sci Monit In Press; DOI: 10.12659/MSM.943738
07 Mar 2024 : Clinical Research
Knowledge of and Attitudes Toward Clinical Trials: A Questionnaire-Based Study of 179 Male Third- and Fourt...Med Sci Monit In Press; DOI: 10.12659/MSM.943468
08 Mar 2024 : Animal Research
Modification of Experimental Model of Necrotizing Enterocolitis (NEC) in Rat Pups by Single Exposure to Hyp...Med Sci Monit In Press; DOI: 10.12659/MSM.943443
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952