31 August 2020: Database Analysis
Identification of Prognostic Biomarkers Associated with Cancer Stem Cell Features in Prostate Adenocarcinoma
Dehua Zhang1ACD, Muchun Zhang2B, Qifu Zhang3DF, Zhiyi Zhao3BCF, Yong Nie1ADE*DOI: 10.12659/MSM.924543
Med Sci Monit 2020; 26:e924543
Abstract
BACKGROUND: Prostate adenocarcinoma (PRAD) is the second most common malignancy in males and the fifth leading cause of cancer mortality. Cancer stem cells (CSCs) play an important role in the occurrence and development of PRAD, however, the prognostic biomarkers associated with CSC features have not been identified in PRAD.
MATERIAL AND METHODS: In order to identify the prognostic stemness-related genes (SRGs) of PRAD, the RNA sequencing data of patients with PRAD were retrieved from The Cancer Genome Atlas (TCGA) databases. The mRNA expression-based stemness index (mRNAsi) and the differential expressed genes (DEGs) were evaluated and identified. The associations between the mRNAsi and tumorigenesis, overall survival (OS), prostate-specific antigen (PSA) value, and Gleason score were also established by nonparametric test and Kaplan-Meier survival analysis. The SRGs were identified as the overlapped DEGs of PRAD-associated DEGs and the mRNAsi-associated DEGs. Based on the prognostic SRGs, the predict model was constructed. Its accuracy was tested by the area under the curve (AUC) of the receiver operator characteristic (ROC) curve and the risk score.
RESULTS: A total of 6005 PRAD-associated DEGs and 2462 mRNAsi-associated DEGs were identified. The mRNAsi was significantly upregulated in PRAD and associated with the PSA value and Gleason score. A total of 1631 SRGs were identified, with 36 prognostic SRGs screened by the univariate Cox analysis. Based on the prognostic SRGs, the predict model was constructed with the AUC of 0.986. Moreover, the risk score of the model was proved to be an independent prognostic factor, indicating its significant applicability.
CONCLUSIONS: Our data demonstrate the mRNAsi as a reliable index for the tumorigenesis, PSA value, and Gleason score of PRAD. Additionally, this study provides a well-applied model for predicting the OS for patients with PRAD based on prognostic SRGs.
Keywords: carcinogenesis, Decision Support Techniques, Neoplastic Stem Cells, Prostatic Neoplasms, Area Under Curve, Atlases as Topic, Biomarkers, Tumor, DNA Methylation, Databases, Genetic, Gene Expression Profiling, Gene Regulatory Networks, Neoplasm Grading, Prostate-Specific Antigen, ROC Curve
Background
Prostate adenocarcinoma (PRAD) is the second most common malignancy in males and the fifth leading cause of cancer mortality [1,2]. Generally, PRAD is clinically localized, and radical prostatectomy (RP) is the standard treatment option for these patients [3]. RP provides adequate local control and a favorable prognosis [4]. However, many cases experience tumor recurrence and metastasis, in especial bone metastasis [5]. As for patients with advanced PRAD, androgen deprivation therapy (ADT) is usually used, however, these patients also have poor clinical outcomes [6]. Thus, it is crucial to investigate PRAD occurrence and progression factors and to identify potential biomarkers for predicting the prognosis of patients with PRAD.
Like many solid tumors, PRAD is heterogeneous, including many types of cancer cells [7]. Generally, the differentiation of tumor cells is initiated from a population of cancer stem cells (CSCs). CSCs exhibit similar phenotypic and functional properties of normal stem cells, such as self-renewal and reestablishing the heterogeneous tumor cell population [8]. Thus, CSCs are significantly associated with the occurrence, development, and treatment resistance of PRAD [9]. Identification of CSC-associated biomarkers may predict the tumorigenesis and prognosis of PRAD and provide CSC-based diagnostic and therapeutic strategies [10].
Nowadays, CSC features have been explored by deep learning methods to assess the degree of oncogenic dedifferentiation [11]. The DNA methylation-based stemness index (mDNAsi) is used to reflect epigenetic features and the other mRNA expression-based stemness index (mRNAsi) is reflective of gene expression. Both stemness indices can be related to the activity of CSCs, tumor dedifferentiation, and pathological grade. However, the applicability of the mRNAsi in predicting the tumorigenesis and prognosis of PRAD has not been verified.
In this study, RNA-seq data and clinical information of PRAD samples were collected from The Cancer Genome Atlas (TCGA) databases. Additionally, we identified the mRNAsi and the differential expressed genes (DEGs). The stemness-related genes (SRGs) were identified as the overlapped DEGs of PRAD-associated DEGs and the mRNAsi-associated DEGs. Based on the prognostic SRGs, the predict model was constructed. Thus, this study was the first one focused on the role of the mRNAsi in the PRAD, and to suggest that the mRNAsi could be a reliable prognostic index for PRAD patients which may assist oncologists in clinical diagnosis and treatment.
Material and Methods
DATA ACQUISITION:
This study was approved by the Ethics Committee of the Yiling Hospital of Yichang (No. LW2020-04-1401). The gene expression profiling of 498 primary PRAD and 52 normal solid tissue available from the TCGA database (
ESTIMATION OF THE MRNASI BASED ON THE GENE EXPRESSION PROFILING:
The mRNAsi have been reported by Tathiane et al., using an algorithm named one-class logistic regression machine learning (OCLR) to define stem cell signatures of the bulk tissue (CSCs) [11]. In this study, the mRNAsi of PRAD and normal solid tissue were estimated by OCLR based on the normalized gene expression profile [log2 (FPKM+1)]. The associations between the mRNAsi and the tumorigenesis, overall survival (OS), prostate-specific antigen (PSA) value, and Gleason score were also established by nonparametric test and Kaplan-Meier survival analysis.
IDENTIFICATION OF DEGS:
EdgeR method was used to identify the DEGs between 498 primary PRAD and 52 normal solid tissue. The gene with log2 fold change (FC) >1.0 or <−1.0, and false discovery rate (FDR) value <0.05 was defined as DEG. Besides, the PRAD samples where the mRNAsi was greater than the median was defined as the high stemness tumor and vice versa (the low stemness tumor). Similarly, DEGs analysis between low and high stemness PRAD was also conducted. Furthermore, the Gene Oncology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) functional enrichment analysis were utilized to explore the biological processes and pathways that DEGs enriched.
CONSTRUCTION OF MULTIVARIATE COX MODEL:
The SRGs were identified as the overlapped DEGs of PRAD-associated DEGs and the mRNAsi-associated DEGs. The prognostic SRGs selected by univariate Cox analysis were integrated into multivariate model. The accuracy of the model was tested by the area under the curve (AUC) of the receiver operator characteristic (ROC) curve. A protein-protein interaction (PPI) network were constructed based on prognostic genes in multivariate model by STRING database [12].
INDEPENDENT PROGNOSIS ANALYSIS:
For each PRAD sample, the risk score was calculated by the formula generated by the multivariate Cox model as followed:
For each single PRAD sample, “n” represented the number of prognostic gene in the multivariate model; “x” represented the number of each patient; and “β” represented coefficient of each prognostic gene. Additionally, the patients were divided into high and low risk group by the median of risk score. Then, the Kaplan-Meier survival analysis was applied to predict the prognosis value of the risk group. Besides, the independent prognosis value for the risk score was evaluated by univariate and multivariate Cox regression analysis, corrected by demographics, Gleason score and new tumor event during the follow-up period.
STATISTICS ANALYSIS:
As for descriptive statistics, mean±standard deviation was used for the continuous variables in normal distribution while the mean (range) was used for continuous variables in abnormal distribution. In addition, categorical variables were described by counts and percentages. The ROC curve illustrated the sensitivity and specificity of multivariate models to predict 5-year overall mortality. R software (Institute for Statistics and Mathematics, Vienna, Austria;
Results
DEGS IDENTIFICATION:
The baseline clinical characteristics of all patients with PRAD are summarized in Table 1 and the flowchart of this study is shown in Figure 1. The essential characteristics of the 498 primary PRAD and 52 normal solid tissue with RNA-seq data are summarized in Table 2.
A total of 6005 genes including 2697 downregulated genes and 3308 upregulated genes were identified as DEGs between 498 primary PRAD and 52 normal solid tissue. The heatmap and volcano plot were presented in Figure 2A and 2B. The DEGs enriched in the GO term pointed to muscle system process, collagen-containing extracellular matrix, and passive transmembrane transporter activity (Figure 2C), while those enriched in KEGG term were associated with neuroactive ligand-receptor interaction (Figure 2D). Similarly, 2462 genes including 2029 downregulated genes and 433 upregulated genes were identified as DEGs between low and high mRNAsi PRAD. The heatmap and volcano plot are described in Figure 3A and 3B. The stemness-related DEGs enriched in GO (Figure 3C) and KEGG (Figure 3D) terms also included muscle system process, collagen-containing extracellular matrix, passive transmembrane transporter activity and neuroactive ligand-receptor interaction.
IDENTIFICATION OF SRGS WITH PROGNOSTIC VALUE:
As shown in Venn plot, 1631 genes between the PRAD-associated DEGs and 2462 the mRNAsi-associated DEGs were identified as SRGs and included in the univariate Cox analysis (Figure 4A). A total of 36 SRGs with prognostic value were identified by the univariate Cox analysis (Figure 4B). Besides, the mRNAsi of PRAD tissues was significantly higher than that in the normal tissues (Figure 4C, P<0.001). Additionally, the mRNAsi was also significantly associated with the Gleason score (Figure 4D, P<0.001) and PSA value (Figure 4E, P<0.001). Although patients with high mRNAsi had a decreased OS compared to patients with low mRNAsi, the difference was not significant (Figure 4F, P=0.059).
The prognostic SRGs were then integrated into the multivariate Cox model. The risk score distribution based on risk score of each sample was described in the scatter plot (Figure 5A) and risk curve (Figure 5B). The risk score was demonstrated to be a prognostic factor for OS in the Kaplan-Meier analysis (Figure 5C, P<0.001). In addition, the efficiency and residual distribution of the multivariate model was accessed by the ROC curve (AUC=0.986) and residual plot (Figure 5D, 5E).
INDEPENDENT PROGNOSIS ANALYSIS AND CONSTRUCTION OF PPI NETWORK:
The risk score was also revealed to be an independently prognostic indicator for patients with PRAD in univariate (hazard ratio [HR]=1.573, 95% confidence interval [CI] (1.551–1.595), P<0.001) (Figure 6A) and multivariate (HR=1.343, 95% CI (1.323–1.364), <0.001) (Figure 6B) Cox regression model was corrected by demographics, Gleason score, and new tumor event during the follow-up period.
A PPI network were also constructed based on prognostic SRGs in multivariate model by STRING database to present their interaction. In Figure 7, prognostic SRGs, such as KIF18B, KIF4A, BIRC5, NCAPG, UBE2C, NEIL3, and EXO1, had a strong interaction relationship.
Discussion
PRAD is the second most challenging cause of cancer-related deaths in men [1,13] and its mortality is mainly associated with tumor progression [13]. CSCs play an important role in the occurrence and progression of PRAD and therapy resistance during treatment [9,14]. Thus, there is a pressing need to explore the prognostic SRGs which may assist oncologists in early diagnosis and targeted therapy. In this study, our findings firstly demonstrated that the mRNAsi was a reliable index which was significantly upregulated in PRAD. Based on the prognostic SRGs, we constructed a well-applied prediction model to predict the prognosis of PRAD patients and identified the interaction network of key prognostic SRGs.
CSCs possess the self-renewal properties, genomic instability, and are regarded as the engine of tumor evolution. In tumor metabolism of PRAD, CSCs regulate the MYC-dependent metabolic reprogramming by increasing the expression levels of GLS1 glutaminase to utilize the glutamine [15,16]. The glutamine utilization is also mediated by the androgen receptor (AR), whose gene amplification or mutation works in the “androgen-independent” tumor progression [17]. Thus, the CSC-mediated androgen-resistance is associated with AR and glutamine metabolism [18]. Additionally, CSCs can also induce the epithelial-mesenchymal transition (EMT) in PRAD which is associated with the tumor metastasis [19].
As the important roles of CSCs in the tumor initiating, tumor progression, and drug resistance of PRAD become known, identification of CSC-related signatures can be used to monitor the tumor development, recurrence, and metastasis; predict the prognosis of patients; and evaluate the efficacy of therapeutic methods. In this study, we used the mRNAsi as the stemness index of transcriptome gene expression and found that the mRNAsi was significantly highly expressed in the PRAD tissue samples compared with normal tissue [20]. Although the patients with higher mRNAsi had a lower OS, due to the favorable prognosis of patients with PRAD, the difference was not significant (
It has been previously reported that the mRNAsi was related to the biological processes which were active in CSCs and malignant histopathological grade [11]. Besides, advanced neoplasms were also believed to have high stemness index (mRNAsi), because of the aggressiveness nature and dedifferentiated phenotype of the metastatic and recurrent tumor cells [21]. Moreover, the mRNAsi was also found to be related to some novel oncogenic pathways and somatic alterations, which demonstrate the roles of CSC in the tumorigenesis [11].
Screening of serum PSA is the conventional and convenient method for predicting the probability of PCa and widely used in the worldwide [22]. However, its benefit is still controversial, especially in the older men (≥70 or 75 years) or patients with limited life expectancy (<10 years) [23]. Gleason score is also a reliable grading system for the aggressiveness for PRAD and strongly predicts the prognosis of patients with PRAD [24]. The high-grade PRAD can be defined as Gleason score sum 7 or higher [22]. The evaluation of Gleason grading in predicting PRAD behavior and recurrence is similar to expert pathologists by the deep-learning system [25]. However, the effects of PSA and Gleason score in PRAD prognosis are still queried by many scientists and exploring the novel prognostic biomarkers is an urgent issue [26]. In this study, we found the stemness index mRNAsi was also significantly associated with the PSA value and Gleason score, which may provide a reliable index for the prognosis of PRAD.
In view of the important roles of CSCs in the tumorigenesis, progression, and drug resistance, we identified the key prognostic SRGs, such as KIF18B, KIF4A, BIRC5, NCAPG, UBE2C, NEIL3, and EXO1. Based on the prognostic SRGs, we built a model to predict the OS of patients with PRAD and the model achieved a good accuracy and applicability (AUC: 0.986). Evaluating the prognostic factors is important for oncologists in clinical decision-marking, thus many previous studies have focused on the identification of predictors for patients with PRAD [5,27,28]. Clinical information (such as age), blood examination (such as PSA value), tumor characteristics (such as androgen-independent and androgen-dependent type) and treatment methods (such as RP and ADT) have been regarded as prognostic factors and proved to be associated with the prognosis of patients with PRAD [28,29]. However, these clinical factors do not include the molecular biomarkers. With the well-application of TCGA dataset, regulatory network of miRNA, mRNA, and alternative splicing has been constructed and prognosis-associated biomarkers have also been identified for patients with PRAD [31,32]. Unfortunately, these prediction model do not cover the CSC-related signatures and prognostic SRGs. Thus, our study is an appropriate supplement to the existing prediction models for the prognosis evaluation of PRAD patients.
Although this study was the first one which suggested good accuracy of the mRNAsi in predicting the prognosis of PRAD and constructed a well-applied prediction model based on the CSC-related genes, it still possessed some shortcomings that warrant consideration. Firstly, all the samples involved in this study are from America, and thus we are not quite sure about the applicability of prediction model in European and Asian. Second, this proposed prediction model has not been verified by external databases. Third, the protein expression of key prognostic SRGs have not been verified. Thus, our next study will focus on validating the prediction model and key prognostic SRGs using our own data and other public database.
Conclusions
Our data demonstrated that the mRNAsi as a reliable index for tumorigenesis, PSA value, and Gleason score of PRAD. Additionally, this study provided a well-applied model for predicting the OS for patients with PRAD based on prognostic SRGs.
Figures
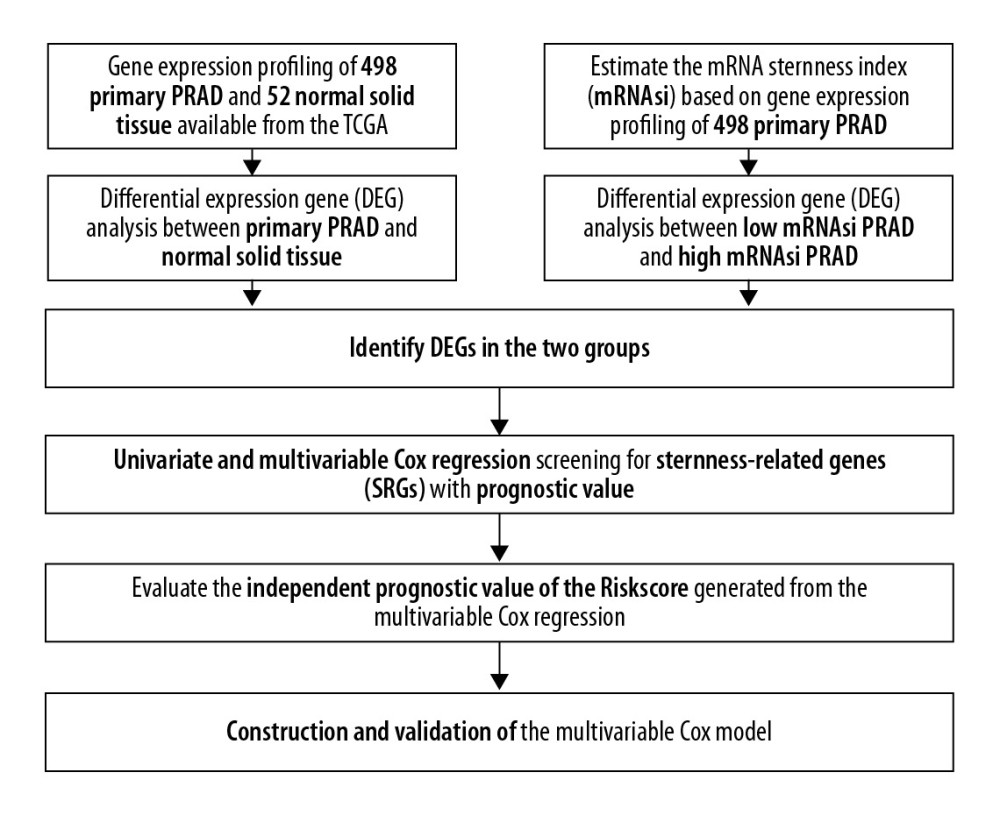 Figure 1. The flowchart of all analysis process.
Figure 1. The flowchart of all analysis process. 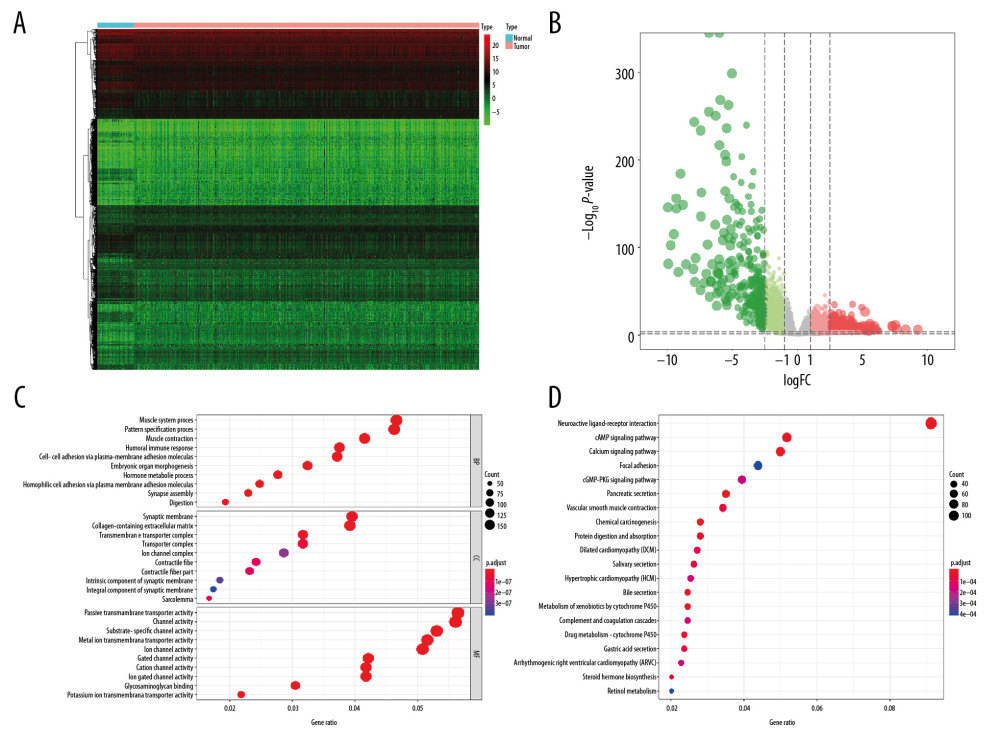 Figure 2. The results of differential expression genes analysis and functional enrichment analysis between 498 primary PRAD and 52 normal solid tissue. A total of 6005 genes including 2697 downregulated ones and 3308 upregulated ones were identified as DEGs between 498 primary PRAD and 52 normal solid tissue, illustrating in heatmap (A) and volcano plot (B). Most significant GO (C) and KEGG (D) terms for DEG were muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity, and neuroactive ligand–receptor interaction, illustrating in bubble plot. PRAD – prostate adenocarcinoma; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes.
Figure 2. The results of differential expression genes analysis and functional enrichment analysis between 498 primary PRAD and 52 normal solid tissue. A total of 6005 genes including 2697 downregulated ones and 3308 upregulated ones were identified as DEGs between 498 primary PRAD and 52 normal solid tissue, illustrating in heatmap (A) and volcano plot (B). Most significant GO (C) and KEGG (D) terms for DEG were muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity, and neuroactive ligand–receptor interaction, illustrating in bubble plot. PRAD – prostate adenocarcinoma; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes. 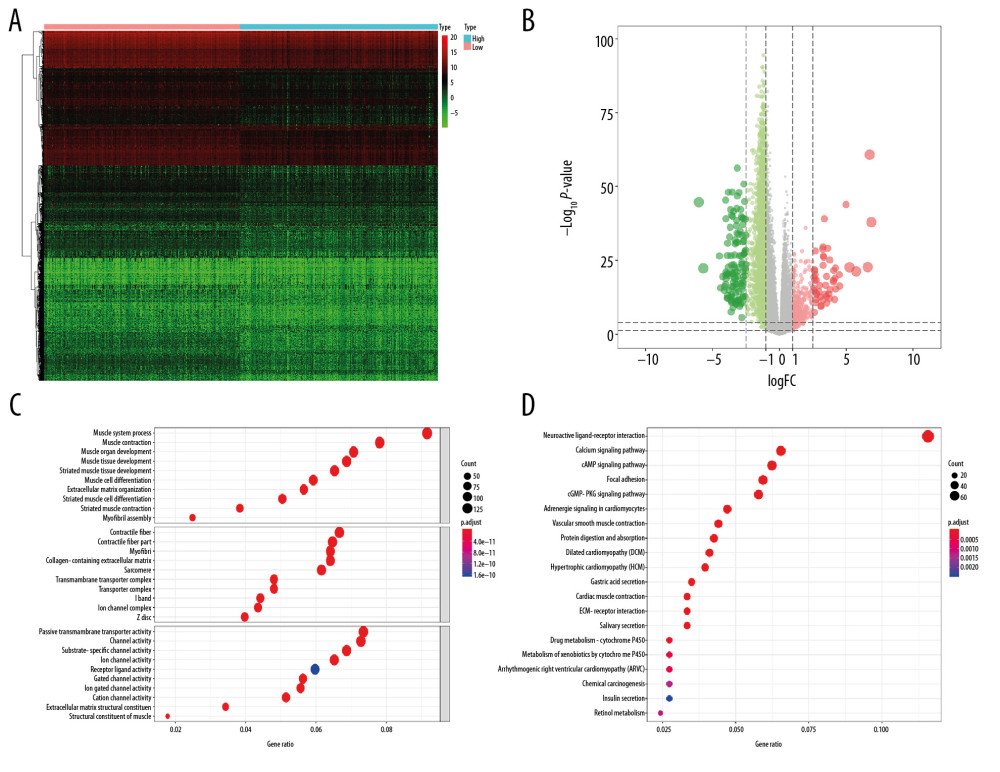 Figure 3. The results of differential expression genes analysis and functional enrichment analysis between low mRNAsi PRAD and high mRNAsi PRAD. A total of 2462 genes including 2029 downregulated ones and 433 upregulated ones were identified as DEGs between low mRNAsi PRAD and high mRNAsi PRAD, illustrating in heatmap (A) and volcano plot (B). GO (C) and KEGG (D) terms including muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity and neuroactive ligand–receptor interaction were identified as significantly enriched items in stemness-related DEGs, illustrating in bubble plot. PRAD – prostate adenocarcinoma; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes.
Figure 3. The results of differential expression genes analysis and functional enrichment analysis between low mRNAsi PRAD and high mRNAsi PRAD. A total of 2462 genes including 2029 downregulated ones and 433 upregulated ones were identified as DEGs between low mRNAsi PRAD and high mRNAsi PRAD, illustrating in heatmap (A) and volcano plot (B). GO (C) and KEGG (D) terms including muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity and neuroactive ligand–receptor interaction were identified as significantly enriched items in stemness-related DEGs, illustrating in bubble plot. PRAD – prostate adenocarcinoma; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes. 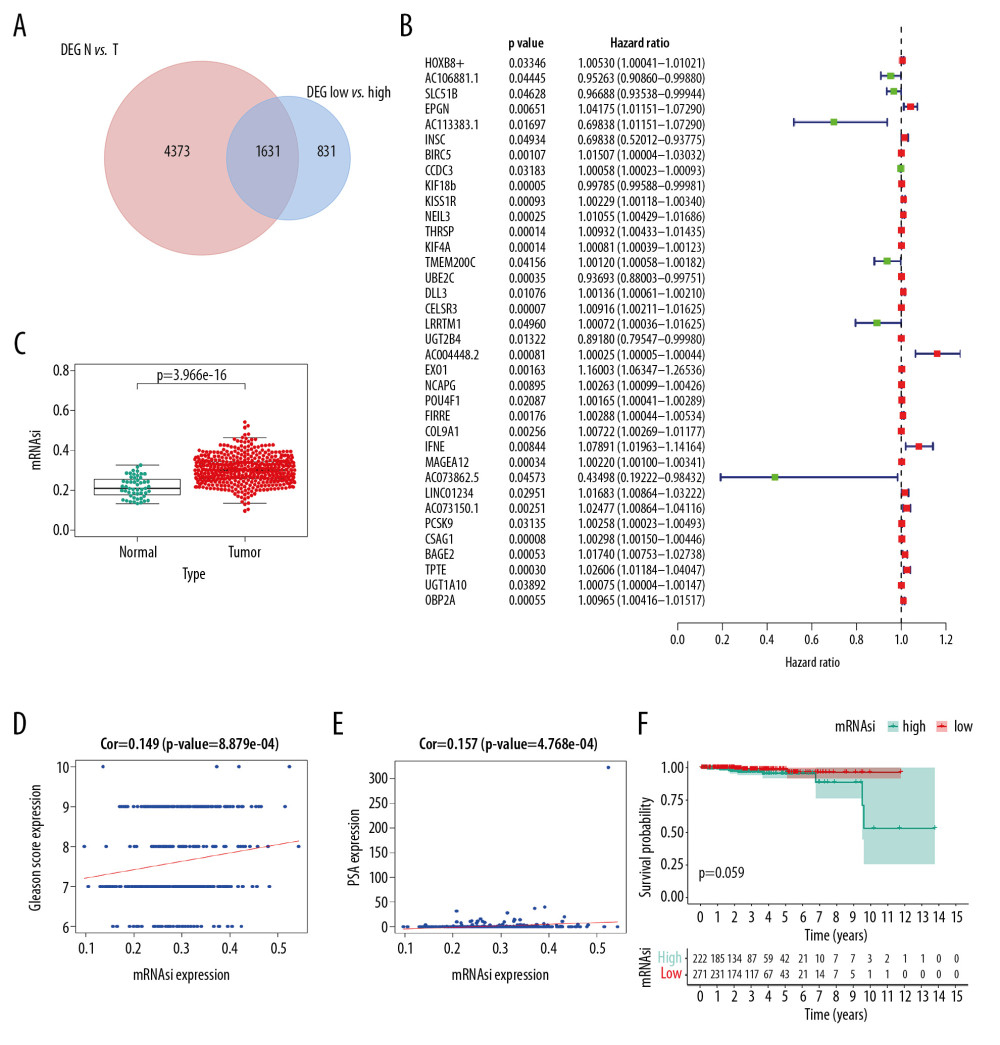 Figure 4. Identification of SRGs with prognostic value. As shown in Venn plot, 1631 DEGs between both high/low stemness tumors and tumor/normal tissues were screened as SRGs and included in the univariate Cox analysis (A). The results of univariate Cox analysis of SRGs with prognostic value were illustrated in the forest plot (B). Besides, significant difference in mRNAsi between normal and tumor tissues was identified (C, P<0.001). And mRNAsi of PRAD were associated with Gleason score (D, P<0.001), PSA value (E, P<0.001) and prognosis (F, P=0.059). SRGs – stemness-related genes; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; PRAD – prostate adenocarcinoma; PSA – prostate-specific antigen.
Figure 4. Identification of SRGs with prognostic value. As shown in Venn plot, 1631 DEGs between both high/low stemness tumors and tumor/normal tissues were screened as SRGs and included in the univariate Cox analysis (A). The results of univariate Cox analysis of SRGs with prognostic value were illustrated in the forest plot (B). Besides, significant difference in mRNAsi between normal and tumor tissues was identified (C, P<0.001). And mRNAsi of PRAD were associated with Gleason score (D, P<0.001), PSA value (E, P<0.001) and prognosis (F, P=0.059). SRGs – stemness-related genes; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; PRAD – prostate adenocarcinoma; PSA – prostate-specific antigen. 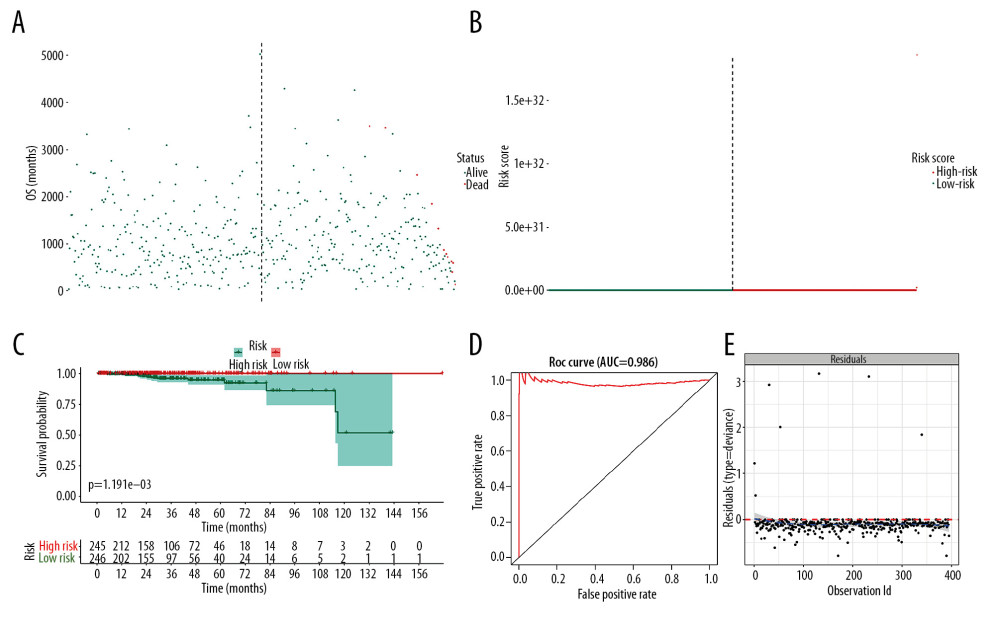 Figure 5. The model diagnosis results of multivariate Cox model constructed based on prognostic SRGs. Prognostic SRGs were integrated into multivariate Cox model. The scatter plot (A) and risk curve (B) of the model demonstrated the risk score distribution based on risk score of each sample. Kaplan-Meier curve suggested the prognostic value of the risk score (C, P<0.001). And the efficiency and residual distribution of the multivariate model was accessed by the ROC curve (AUC=0.860) and residual plot (D, E). SRGs – stemness-related genes; AUC – area under the curve; ROC – receiver operator characteristic.
Figure 5. The model diagnosis results of multivariate Cox model constructed based on prognostic SRGs. Prognostic SRGs were integrated into multivariate Cox model. The scatter plot (A) and risk curve (B) of the model demonstrated the risk score distribution based on risk score of each sample. Kaplan-Meier curve suggested the prognostic value of the risk score (C, P<0.001). And the efficiency and residual distribution of the multivariate model was accessed by the ROC curve (AUC=0.860) and residual plot (D, E). SRGs – stemness-related genes; AUC – area under the curve; ROC – receiver operator characteristic. 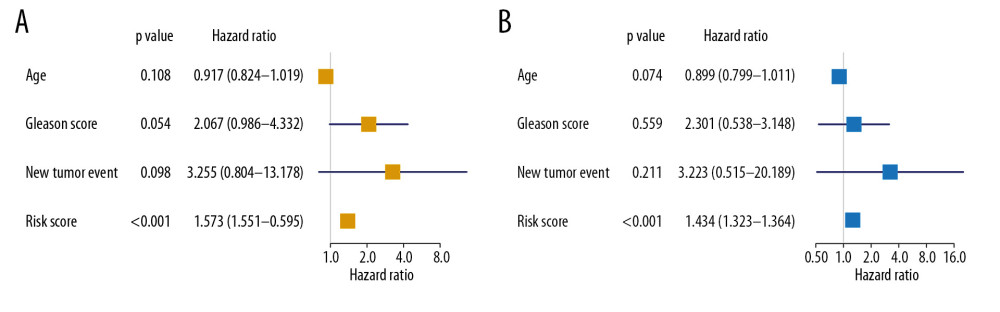 Figure 6. Independent prognosis analysis of the risk score generated from the multivariate Cox model including prognostic SRGs. In univariate (HR=1.573, 95% CI (1.551–1.595), P<0.001) (A) and multivariate (HR=1.343, 95% CI (1.323–1.364), P<0.001) (B) Cox regression model corrected by demographics, Gleason score and new tumor event during the follow-up period, the risk score was shown to be an independently prognostic indicator for PRAD patients. SRGs – stemness-related genes; HR – hazard ratio; CI – confidence interval; PRAD – prostate adenocarcinoma.
Figure 6. Independent prognosis analysis of the risk score generated from the multivariate Cox model including prognostic SRGs. In univariate (HR=1.573, 95% CI (1.551–1.595), P<0.001) (A) and multivariate (HR=1.343, 95% CI (1.323–1.364), P<0.001) (B) Cox regression model corrected by demographics, Gleason score and new tumor event during the follow-up period, the risk score was shown to be an independently prognostic indicator for PRAD patients. SRGs – stemness-related genes; HR – hazard ratio; CI – confidence interval; PRAD – prostate adenocarcinoma. 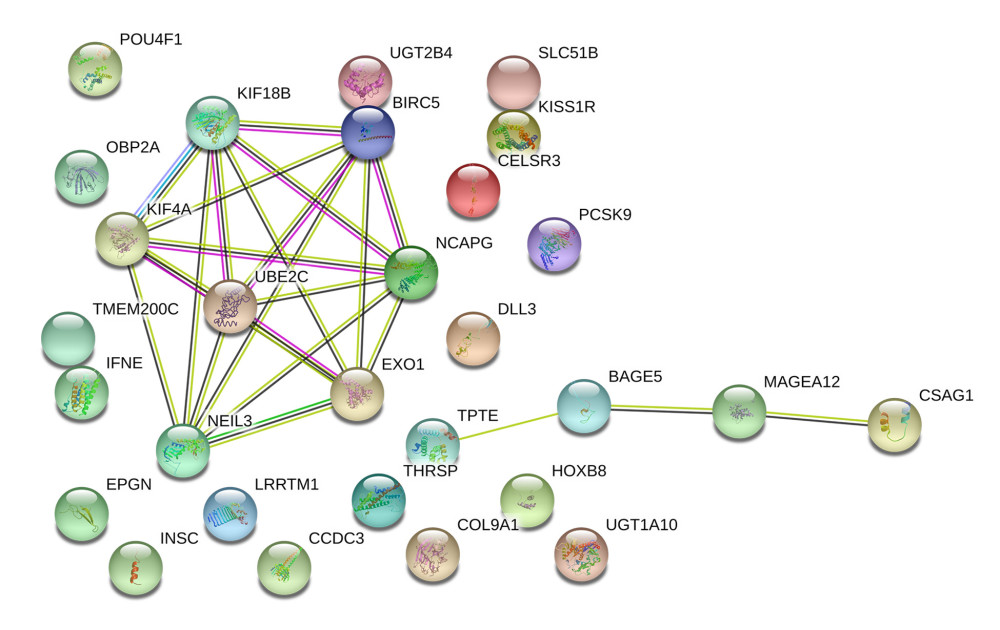 Figure 7. The construction of the PPI network based on prognostic SRGs. PPI – protein–protein interaction; SRGs – stemness-related genes.
Figure 7. The construction of the PPI network based on prognostic SRGs. PPI – protein–protein interaction; SRGs – stemness-related genes. References
1. Siegel R, Naishadham D, Jemal A, Cancer statistics, 2013: Cancer J Clin, 2013; 63(1); 11-30
2. Ferlay J, Soerjomataram I, Dikshit R, Cancer incidence and mortality worldwide: Sources, methods and major patterns in GLOBOCAN 2012: Int J Cancer, 2015; 136(5); E359-86
3. van den Bergh RC, Giannarini G, Prostate cancer: Surgery versus observation for localized prostate cancer: Nat Rev Urol, 2014; 11(6); 312-13
4. Hoffman RM, Shi Y, Freedland SJ, Treatment patterns for older veterans with localized prostate cancer: Cancer Epidemiol, 2015; 39(5); 769-77
5. Meng T, Chen R, Zhong N, Factors associated with improved survival following surgical treatment for metastatic prostate cancer in the spine: Retrospective analysis of 29 patients in a single center: World J Surg Oncol, 2016; 14(1); 200
6. Boukovala M, Spetsieris N, Weldon JA, A candidate androgen signalling signature predictive of response to abiraterone acetate in men with metastatic castration-resistant prostate cancer: Eur J Cancer, 2020; 127; 67-75
7. Packer JR, Maitland NJ, The molecular and cellular origin of human prostate cancer: Biochim Biophys Acta, 2016; 1863(6 Pt A); 1238-60
8. Leao R, Domingos C, Figueiredo A, Cancer stem cells in prostate cancer: implications for targeted therapy: Urol Int, 2017; 99(2); 125-36
9. Skvortsov S, Skvortsova , Tang DG, Dubrovska A, Concise review: Prostate cancer stem cells: current understanding: Stem Cells, 2018; 36(10); 1457-74
10. Passaniti A, Hussain A, Novel approaches targeting mitochondrial fission to deplete stem-like tumor cells in prostate cancer and improve treatment outcomes: Ann Transl Med, 2019; 7(Suppl 8); S335
11. Malta TM, Sokolov A, Gentles AJ, Machine learning identifies stemness features associated with oncogenic dedifferentiation: Cell, 2018; 173(2); 338-54.e315
12. Szklarczyk D, Franceschini A, Wyder S, STRING v10: protein-protein interaction networks, integrated over the tree of life: Nucleic Acids Res, 2015; 43(Database issue); D447-52
13. Siegel R, Ma J, Zou Z, Jemal A, Cancer statistics, 2014: Cancer J Clin, 2014; 64(1); 9-29
14. Gruber M, Handle F, Culig Z, The stem cell inhibitor salinomycin decreases colony formation potential and tumor-initiating population in docetaxel-sensitive and docetaxel-resistant prostate cancer cells: Prostate, 2020; 80(3); 267-73
15. Civenni G, Malek A, Albino D, RNAi-mediated silencing of Myc transcription inhibits stem-like cell maintenance and tumorigenicity in prostate cancer: Cancer Res, 2013; 73(22); 6816-27
16. Dang CV, Le A, Gao P, MYC-induced cancer cell energy metabolism and therapeutic opportunities: Clin Cancer Res, 2009; 15(21); 6479-83
17. Saraon P, Drabovich AP, Jarvi KA, Diamandis EP, Mechanisms of androgen-independent prostate cancer: EJIFCC, 2014; 25(1); 42-54
18. Karacosta LG, Foster BA, Azabdaftari G, A regulatory feedback loop between Ca2+/calmodulin-dependent protein kinase kinase 2 (CaMKK2) and the androgen receptor in prostate cancer progression: J Biol Chem, 2012; 287(29); 24832-43
19. Lo UG, Lee CF, Lee MS, Hsieh JT, The role and mechanism of epithelial-to-mesenchymal transition in prostate cancer progression: Int J Mol Sci, 2017; 18(10) pii: E2079
20. Sokolov A, Paull EO, Stuart JM, One-class detection of cell states in tumor subtypes: Pac Symp Biocomput, 2016; 21; 405-16
21. Pan S, Zhan Y, Chen X, Identification of biomarkers for controlling cancer stem cell characteristics in bladder cancer by network analysis of transcriptome data stemness indices: Front Oncol, 2019; 9; 613
22. Chen R, Xie L, Xue W, Development and external multicenter validation of Chinese Prostate Cancer Consortium prostate cancer risk calculator for initial prostate biopsy: Urol Oncol, 2016; 34(9); 416.e1-7
23. Wilt TJ, Scardino PT, Carlsson SV, Basch E, Prostate-specific antigen screening in prostate cancer: perspectives on the evidence: J Natl Cancer Inst, 2014; 106(3); dju010
24. Dere Y, Celik OI, Celik SY, A grading dilemma; Gleason scoring system: Are we sufficiently compatible? A multi center study: Indian J Pathol Microbiol, 2020; 63(Suppl); S25-29
25. Thomas T, Automated systems comparable to expert pathologists for prostate cancer Gleason grading: Nat Rev Urol, 2020; 17(3); 131
26. Subramanian V, Kattan MW, Prostate cancer: Numeracy and understanding of risk reduction of PSA screening: Nat Rev Urol, 2018; 15(4); 208-9
27. Butler SS, Dee EC, Lamba N, Validation of a subclassification for high-risk prostate cancer in a prospective cohort: Cancer, 2020; 126(10); 2132-38
28. He BM, Chen R, Shi ZK: Front Oncol, 2019; 9; 758
29. Zhou X, Ning Q, Jin K, Development and validation of a preoperative nomogram for predicting survival of patients with locally advanced prostate cancer after radical prostatectomy: BMC Cancer, 2020; 20(1); 97
30. Stangl-Kremser J, Mari A, Suarez-Ibarrola R, Development of a prognostic model for survival time prediction in castration-resistant prostate cancer patients: Urol Oncol; 2020 [Epub ahead of print]
31. Wei J, Yin Y, Deng Q, Integrative analysis of microRNA and gene interactions for revealing candidate signatures in prostate cancer: Front Genet, 2020; 11; 176
32. Cao ZX, Xiao GA, Zhang W, Comprehensive investigation of alternative splicing and development of a prognostic risk score for prostate cancer based on six-gene signatures: J Cancer, 2019; 10(22); 5585-96
Figures
 Figure 1. The flowchart of all analysis process.
Figure 1. The flowchart of all analysis process. Figure 2. The results of differential expression genes analysis and functional enrichment analysis between 498 primary PRAD and 52 normal solid tissue. A total of 6005 genes including 2697 downregulated ones and 3308 upregulated ones were identified as DEGs between 498 primary PRAD and 52 normal solid tissue, illustrating in heatmap (A) and volcano plot (B). Most significant GO (C) and KEGG (D) terms for DEG were muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity, and neuroactive ligand–receptor interaction, illustrating in bubble plot. PRAD – prostate adenocarcinoma; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes.
Figure 2. The results of differential expression genes analysis and functional enrichment analysis between 498 primary PRAD and 52 normal solid tissue. A total of 6005 genes including 2697 downregulated ones and 3308 upregulated ones were identified as DEGs between 498 primary PRAD and 52 normal solid tissue, illustrating in heatmap (A) and volcano plot (B). Most significant GO (C) and KEGG (D) terms for DEG were muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity, and neuroactive ligand–receptor interaction, illustrating in bubble plot. PRAD – prostate adenocarcinoma; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes. Figure 3. The results of differential expression genes analysis and functional enrichment analysis between low mRNAsi PRAD and high mRNAsi PRAD. A total of 2462 genes including 2029 downregulated ones and 433 upregulated ones were identified as DEGs between low mRNAsi PRAD and high mRNAsi PRAD, illustrating in heatmap (A) and volcano plot (B). GO (C) and KEGG (D) terms including muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity and neuroactive ligand–receptor interaction were identified as significantly enriched items in stemness-related DEGs, illustrating in bubble plot. PRAD – prostate adenocarcinoma; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes.
Figure 3. The results of differential expression genes analysis and functional enrichment analysis between low mRNAsi PRAD and high mRNAsi PRAD. A total of 2462 genes including 2029 downregulated ones and 433 upregulated ones were identified as DEGs between low mRNAsi PRAD and high mRNAsi PRAD, illustrating in heatmap (A) and volcano plot (B). GO (C) and KEGG (D) terms including muscle system process, collagen–containing extracellular matrix, passive transmembrane transporter activity and neuroactive ligand–receptor interaction were identified as significantly enriched items in stemness-related DEGs, illustrating in bubble plot. PRAD – prostate adenocarcinoma; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; GO – Gene Ontology; KEGG – Kyoto Encyclopedia of Genes and Genomes. Figure 4. Identification of SRGs with prognostic value. As shown in Venn plot, 1631 DEGs between both high/low stemness tumors and tumor/normal tissues were screened as SRGs and included in the univariate Cox analysis (A). The results of univariate Cox analysis of SRGs with prognostic value were illustrated in the forest plot (B). Besides, significant difference in mRNAsi between normal and tumor tissues was identified (C, P<0.001). And mRNAsi of PRAD were associated with Gleason score (D, P<0.001), PSA value (E, P<0.001) and prognosis (F, P=0.059). SRGs – stemness-related genes; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; PRAD – prostate adenocarcinoma; PSA – prostate-specific antigen.
Figure 4. Identification of SRGs with prognostic value. As shown in Venn plot, 1631 DEGs between both high/low stemness tumors and tumor/normal tissues were screened as SRGs and included in the univariate Cox analysis (A). The results of univariate Cox analysis of SRGs with prognostic value were illustrated in the forest plot (B). Besides, significant difference in mRNAsi between normal and tumor tissues was identified (C, P<0.001). And mRNAsi of PRAD were associated with Gleason score (D, P<0.001), PSA value (E, P<0.001) and prognosis (F, P=0.059). SRGs – stemness-related genes; mRNAsi – mRNA expression-based stemness index; DEGs – differential expressed genes; PRAD – prostate adenocarcinoma; PSA – prostate-specific antigen. Figure 5. The model diagnosis results of multivariate Cox model constructed based on prognostic SRGs. Prognostic SRGs were integrated into multivariate Cox model. The scatter plot (A) and risk curve (B) of the model demonstrated the risk score distribution based on risk score of each sample. Kaplan-Meier curve suggested the prognostic value of the risk score (C, P<0.001). And the efficiency and residual distribution of the multivariate model was accessed by the ROC curve (AUC=0.860) and residual plot (D, E). SRGs – stemness-related genes; AUC – area under the curve; ROC – receiver operator characteristic.
Figure 5. The model diagnosis results of multivariate Cox model constructed based on prognostic SRGs. Prognostic SRGs were integrated into multivariate Cox model. The scatter plot (A) and risk curve (B) of the model demonstrated the risk score distribution based on risk score of each sample. Kaplan-Meier curve suggested the prognostic value of the risk score (C, P<0.001). And the efficiency and residual distribution of the multivariate model was accessed by the ROC curve (AUC=0.860) and residual plot (D, E). SRGs – stemness-related genes; AUC – area under the curve; ROC – receiver operator characteristic. Figure 6. Independent prognosis analysis of the risk score generated from the multivariate Cox model including prognostic SRGs. In univariate (HR=1.573, 95% CI (1.551–1.595), P<0.001) (A) and multivariate (HR=1.343, 95% CI (1.323–1.364), P<0.001) (B) Cox regression model corrected by demographics, Gleason score and new tumor event during the follow-up period, the risk score was shown to be an independently prognostic indicator for PRAD patients. SRGs – stemness-related genes; HR – hazard ratio; CI – confidence interval; PRAD – prostate adenocarcinoma.
Figure 6. Independent prognosis analysis of the risk score generated from the multivariate Cox model including prognostic SRGs. In univariate (HR=1.573, 95% CI (1.551–1.595), P<0.001) (A) and multivariate (HR=1.343, 95% CI (1.323–1.364), P<0.001) (B) Cox regression model corrected by demographics, Gleason score and new tumor event during the follow-up period, the risk score was shown to be an independently prognostic indicator for PRAD patients. SRGs – stemness-related genes; HR – hazard ratio; CI – confidence interval; PRAD – prostate adenocarcinoma. Figure 7. The construction of the PPI network based on prognostic SRGs. PPI – protein–protein interaction; SRGs – stemness-related genes.
Figure 7. The construction of the PPI network based on prognostic SRGs. PPI – protein–protein interaction; SRGs – stemness-related genes. Tables
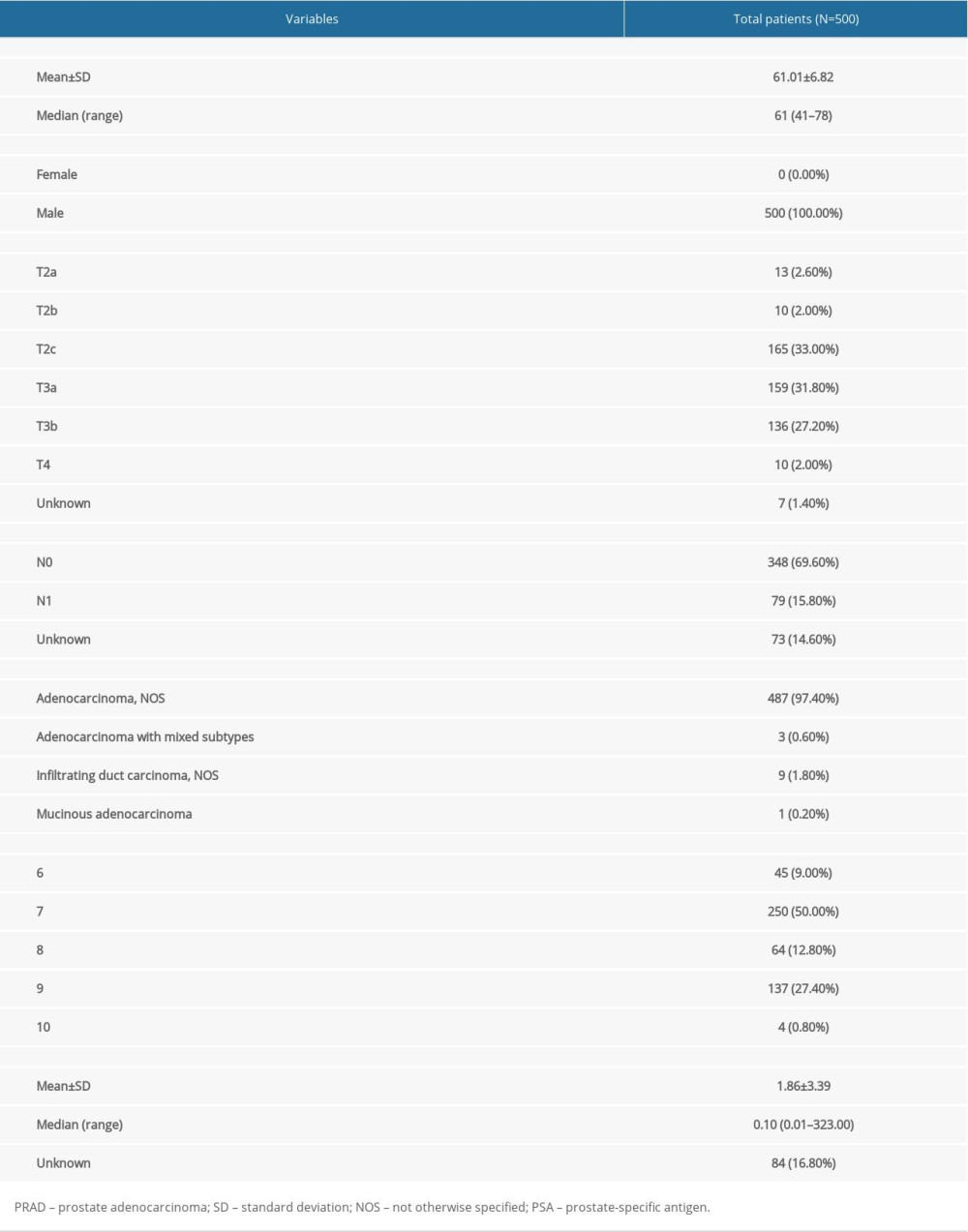 Table 1. Baseline information of patients diagnosed with PRAD.
Table 1. Baseline information of patients diagnosed with PRAD.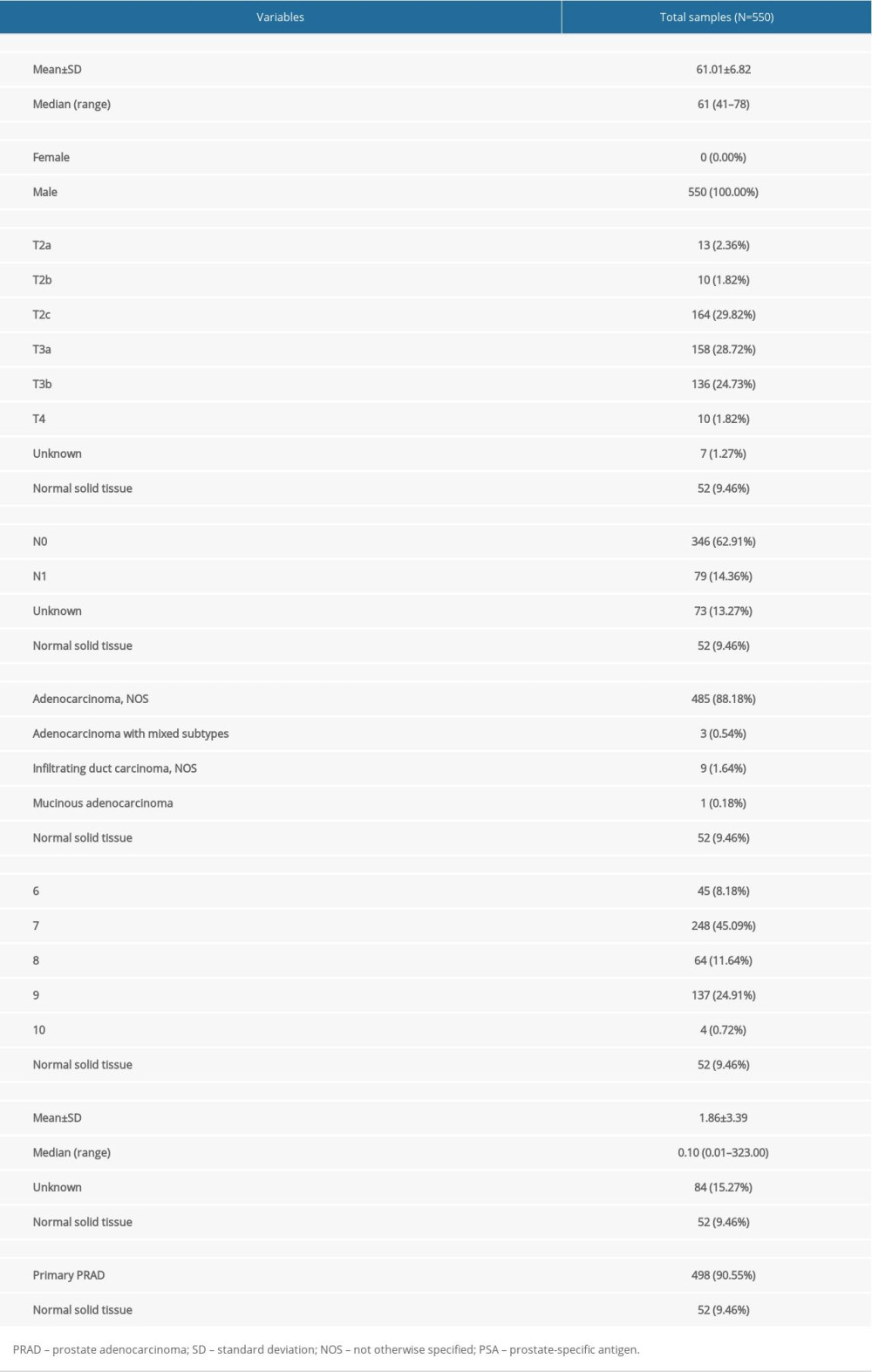 Table 2. Essential characteristics of the primary PRAD and normal solid tissue with RNA-seq data.
Table 2. Essential characteristics of the primary PRAD and normal solid tissue with RNA-seq data. Table 1. Baseline information of patients diagnosed with PRAD.
Table 1. Baseline information of patients diagnosed with PRAD. Table 2. Essential characteristics of the primary PRAD and normal solid tissue with RNA-seq data.
Table 2. Essential characteristics of the primary PRAD and normal solid tissue with RNA-seq data. In Press
21 Mar 2024 : Meta-Analysis
Economic Evaluation of COVID-19 Screening Tests and Surveillance Strategies in Low-Income, Middle-Income, a...Med Sci Monit In Press; DOI: 10.12659/MSM.943863
10 Apr 2024 : Clinical Research
Predicting Acute Cardiovascular Complications in COVID-19: Insights from a Specialized Cardiac Referral Dep...Med Sci Monit In Press; DOI: 10.12659/MSM.942612
06 Mar 2024 : Clinical Research
Enhanced Surgical Outcomes of Popliteal Cyst Excision: A Retrospective Study Comparing Arthroscopic Debride...Med Sci Monit In Press; DOI: 10.12659/MSM.941102
06 Mar 2024 : Clinical Research
Prevalence and Variation of Medical Comorbidities in Oral Surgery Patients: A Retrospective Study at Jazan ...Med Sci Monit In Press; DOI: 10.12659/MSM.943884
Most Viewed Current Articles
17 Jan 2024 : Review article
Vaccination Guidelines for Pregnant Women: Addressing COVID-19 and the Omicron VariantDOI :10.12659/MSM.942799
Med Sci Monit 2024; 30:e942799
14 Dec 2022 : Clinical Research
Prevalence and Variability of Allergen-Specific Immunoglobulin E in Patients with Elevated Tryptase LevelsDOI :10.12659/MSM.937990
Med Sci Monit 2022; 28:e937990
16 May 2023 : Clinical Research
Electrophysiological Testing for an Auditory Processing Disorder and Reading Performance in 54 School Stude...DOI :10.12659/MSM.940387
Med Sci Monit 2023; 29:e940387
01 Jan 2022 : Editorial
Editorial: Current Status of Oral Antiviral Drug Treatments for SARS-CoV-2 Infection in Non-Hospitalized Pa...DOI :10.12659/MSM.935952
Med Sci Monit 2022; 28:e935952








